![PDF] Combined experimental and computational analysis of DNA damage signaling reveals context-dependent roles for Erk in apoptosis and G1/S arrest after genotoxic stress | Semantic Scholar PDF] Combined experimental and computational analysis of DNA damage signaling reveals context-dependent roles for Erk in apoptosis and G1/S arrest after genotoxic stress | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/a9b3aa2cee2d1cfe9693a08279d973a4e6fd09a6/3-Figure1-1.png)
PDF] Combined experimental and computational analysis of DNA damage signaling reveals context-dependent roles for Erk in apoptosis and G1/S arrest after genotoxic stress | Semantic Scholar
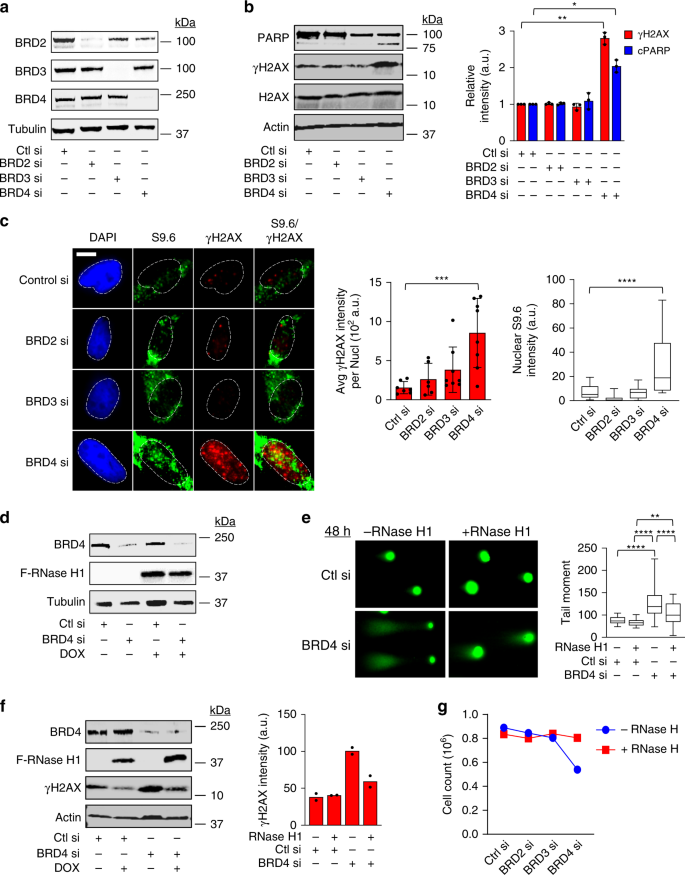
BRD4 prevents the accumulation of R-loops and protects against transcription–replication collision events and DNA damage | Nature Communications
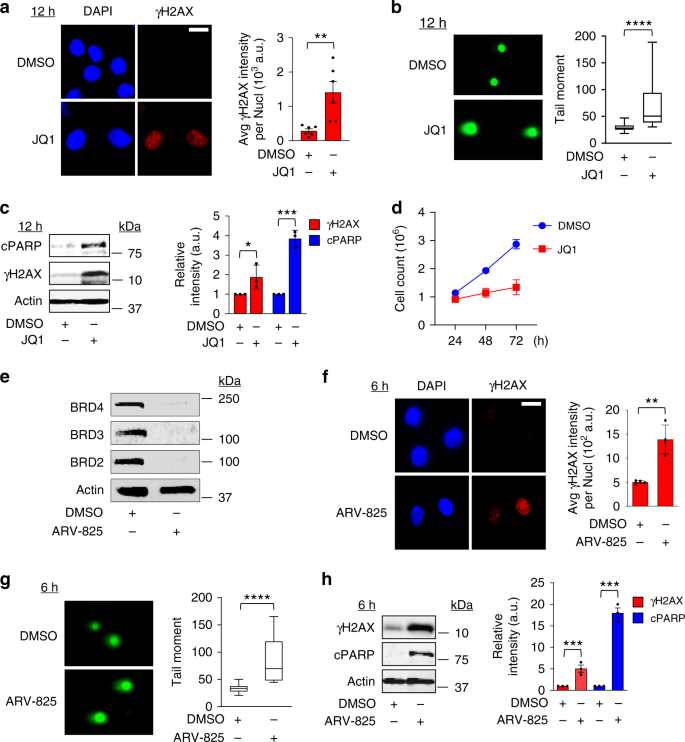
BRD4 prevents the accumulation of R-loops and protects against transcription–replication collision events and DNA damage | Nature Communications
Transite: A Computational Motif-Based Analysis Platform That Identifies RNA-Binding Proteins Modulating Changes in Gene Expressi

Sequential Application of Anticancer Drugs Enhances Cell Death by Rewiring Apoptotic Signaling Networks: Cell
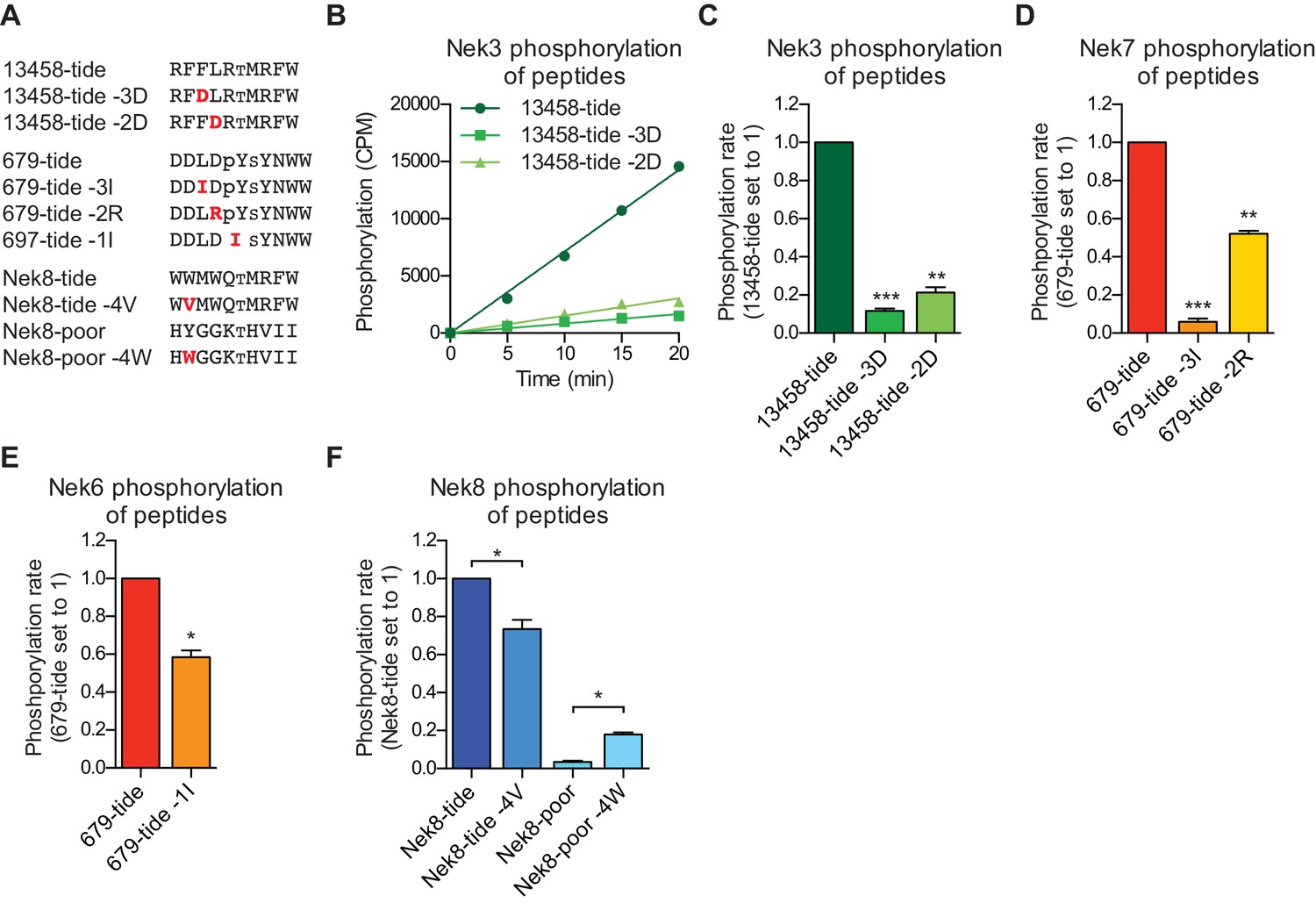
Comprehensive substrate specificity profiling of the human Nek kinome reveals unexpected signaling outputs | eLife

Comprehensive substrate specificity profiling of the human Nek kinome reveals unexpected signaling outputs | eLife
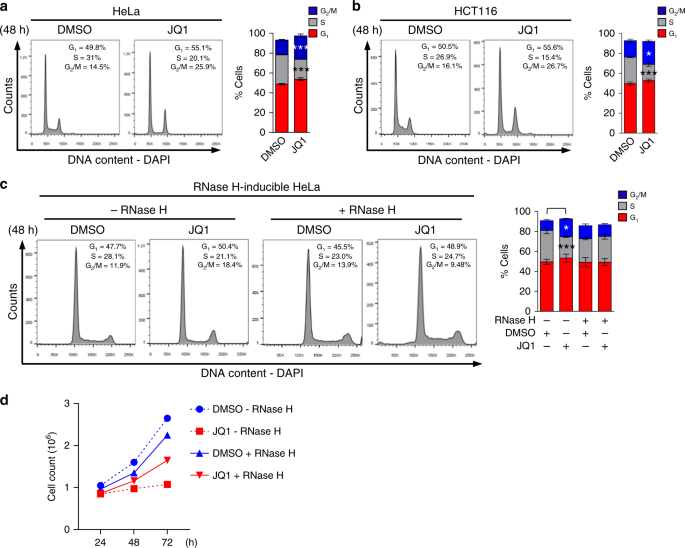
BRD4 prevents the accumulation of R-loops and protects against transcription–replication collision events and DNA damage | Nature Communications

PDF) An integrated comparative phosphoproteomic and bioinformatic approach reveals a novel class of MPM-2 motifs upregulated in EGFRvIII-expressing Glioblastoma Cells
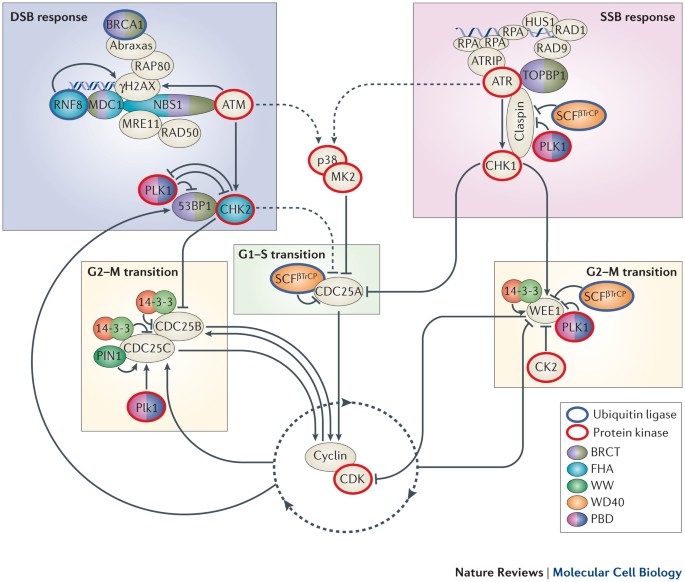
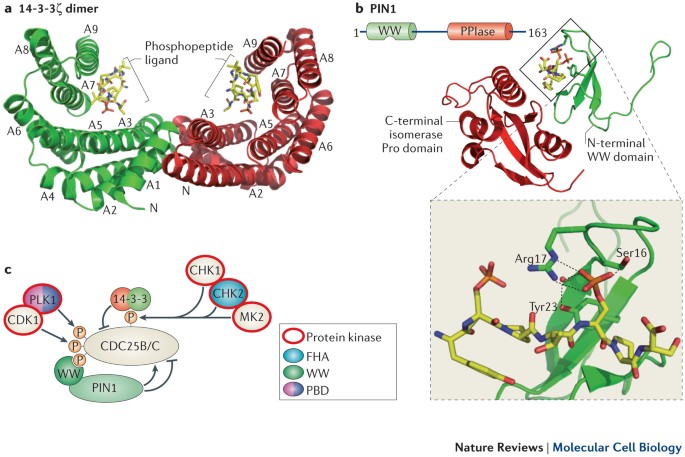
Phospho-Ser/Thr-binding domains: navigating the cell cycle and DNA damage response | Nature Reviews Molecular Cell Biology

PDF) Cytoplasmic Localization of Tristetraprolin Involves 14-3-3-dependent and -independent Mechanisms

PDF) Boyd NF, Martin LJ, Yaffe MJ, Minkin SMammographic density and breast cancer risk: current understanding and future prospects. Breast Cancer Res 13: 223

Phospho-Ser/Thr-binding domains: navigating the cell cycle and DNA damage response | Nature Reviews Molecular Cell Biology

Transite: A Computational Motif-Based Analysis Platform That Identifies RNA-Binding Proteins Modulating Changes in Gene Expression - ScienceDirect
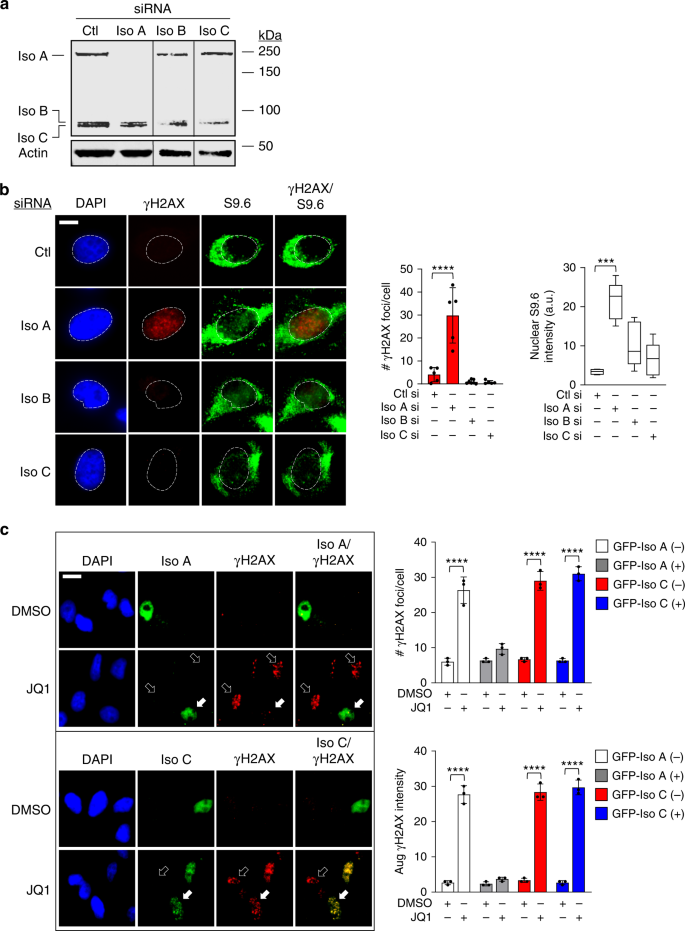
BRD4 prevents the accumulation of R-loops and protects against transcription–replication collision events and DNA damage | Nature Communications

Comprehensive substrate specificity profiling of the human Nek kinome reveals unexpected signaling outputs | eLife